RuleBender:
biological rule-based modeling, simulation, and visualization
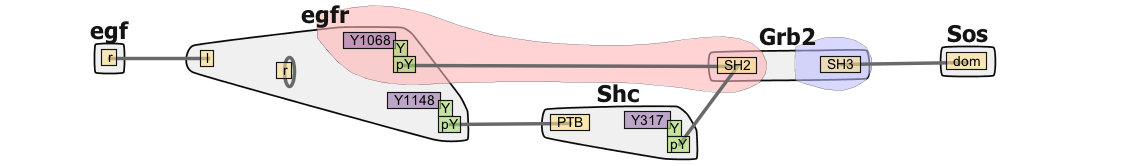
Rule Based Modeling and Simulation with RuleBender
Modern biological experiments generate large amounts of data describing intracellular dynamics. Rule-based languages for describing intracellular biochemistry allow for the construction and simulation of detailed models with unprecedented scope and precision. Rule-Based Models (RBM) can be used to suggest new hypotheses and new ideas for future experimentation.
Rule-based models can be difficult to understand and time consuming to debug. RuleBender is a free tool for constructing, debugging, simulating and analyzing rule-based biological models in the BioNetGen language. The MOSBIE extension further provides a tool for the comparison and analysis of biochemical model families.
When using RuleBender, please cite:
A.M. Smith, W. Xu, Y. Sun, J.R. Faeder, G.E. Marai, “RuleBender: Integrated Modeling, Simulation and Visualization for Rule-Based Intracellular Biochemistry”, BMC Bioinformatics, 13(Suppl 8):S3: 1-24, Jun 2012.
If also using MOSBIE, please cite:
J. Wenskovitch, L.A. Harris, J.J. Tapia, J.R. Faeder, G.E. Marai, "MOSBIE: A Tool for Comparison and Analysis of Rule-Based Biochemical Models", BMC Bioinformatics Journal 15:316, Vol. 15, pp. 1--22, 2014.